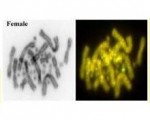
Male or female? In many reptiles sex determination is temperature-dependent. But how this works has been a mystery for the 50 years that has elapsed since the phenomenon was discovered in the African dragon, Agama agama. New insights have emerged from work by Chutian Ge and his collegues who show that ancient conserved epigentic machinary is involved in the thermosensitive regulation of key sex genes. Team Pogona was asked to provide a perspective on the new findings. We have received many enquiries since. Below, we explain more fully what we believe is going on.

Sex determination in reptiles is a complex affair, because incubation temperature and genes interact in many species to regulate sexual development and decide sexual fate, male or female. A central question that has remained unanswered is, what molecular mechanism allows temperature to so profoundly influence the developmental pathways that determine sex?
The means to identify a master sex-determining gene in species with genetic sex determination is well established. First, one must identify genes on the sex chromosomes. Then the challenge becomes one of demonstrating which of these are differentially expressed in male and female embryos early in development. Finally, it is necessary to manipulate the expression of any candidate genes to demonstrate reversal of sex.
The path to identifying the mechanisms of temperature-dependent sex determination (TSD) is not so clearly defined. Temperature could exert its effect on any of the many autosomal genes involved in sexual differentiation, even those peripherally involved, provided their altered expression is capable of reversing sex. For example, thermosensitivity in a gene that encodes for proteins governing in transport of key molecules across the nuclear membrane, potentially isolating those molecules in the cytoplasm at some temperatures but not others, may be the pathway by which temperature exerts its influence of sexual fate.
Once you realise that the above is only one of a very many possibilities that must be disentagled from very many other genes that are differentially expressed in male and female after the decision on sexual fate has been made, the intractability of this problem becomes clear.
With such a wide range of possible candidates for thermal influence, in a sea of other genes that are differntially expressed downstream as a consequence, it is little wonder that, in the 50 years since TSD was discovered in reptiles, we have not advanced far in our understanding of the mechanisms of TSD.
This is about to change. In the May 11 issue of Science, Ge et al. report that transcription of the chromatin modifier gene Kdm6b (=Jmjd3) responds to temperature in the red-eared slider turtle Trachemys scripta elegans, and confers temperature sensitivity to a key sex-determining gene, Dmrt1.
Dmrt1 is a strong candidate for the male sex-determining gene in this TSD species of turtle, consistent with the master sex-determining role of other DM domain–containing genes in some fish, amphibians, and birds. Depending on the species, these DM domain genes initiate and maintain the male sexual trajectory, and suppress genes important for female development during the critical stages of embryogenesis. Presumably, unlike in mammals, Dmrt1 in the turtle is involved in some way in initiating or supporting SOX9 the auto-regulatory positive feedback loop that drives male development, or that it drives transdifferentiation of what would otherwise be a default female pathway.
What Ge et al. (2018) have now demonstrated is a central role in TSD for highly conserved chromatin modifiers including, but not necessarily limited to, KDM6B.
Cell differentiation depends in large part on the selective expression of particular genes, and the timing of that expression. This selective expression of genes is what leads to development of a liver cell lineage rather than a muscle cell lineage, for example. At the organ level, it leads to the development of a testes rather than an ovary. Histone modifiers are responsible for packing of chromatin into chromosomes, and so has influence on the availability of genes and their promoters for transcription, the relative accessability of transcription factors, and to some degree the higher order proximity of genes to distal enhancers. Because the packing of chromatin imposes constraints on cellular processes such as transcription, histone modifiers play an important role in governing the differential expression of genes in different cell lines.
A favourite among epigenetic researchers is trimethylated H3K27 (the trimethylated Lysine 27 on the exposed tail of protein Histone 3). Trimethylated H3K27, written H3K27me3, is a transcriptional repressor that is associated with the promoters of deactivated genes. It acts in opposition to transcriptional activator, such as H3K4me3, and when acetylated rather than trimethylated, becomes a transcriptional activator. H3K27me3 is unusual in that it has only one known methyltransferase, EZH2, that forms part of the Polycomb Repressive Complex 2 (PRC2) responsible for the repression many genes involved in development and cell differentiation.
Ge et al. found that H3K27m3 is associated with the promoter of the sex gene Dmrt1. Presumably in the very early stages of development, when the gonad is undifferentiated, this transcriptional repressor acts in opposition to influential activators (including H3K27me1) to maintain focal cell lineages in a bipotential state. Trimethylated H3K27 was not found on the promoters of any other sex genes that were differentially expressed early in development.
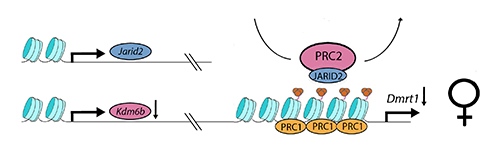
The turtle, Trachemys scripta elegans, has temperature dependent sex determination. At the relatively low temperature of 28C, male offpring are produced; at the relatively high temperature of 32C, female offspring are produced. Ge et al. found that, at the female-producing temperature (see above), the chromatin modifier Kdm6b, a targetted histone demethylase, is downregulated. This presumably allows the PRC2 complex to do its work, through its active component EZH2 and facultative component JARID2 (to form PRC2.2), augmenting and maintaining the trimethylation of H3K27, and enlisting PRC1 to the Dmrt1 promoter to give permanance to the trimethylation and repression through cell division. Dmrt1 repression applies to the focal cell lines, leading to ovary development.
At male-producing temperatures (see below), Kdm6b is upregulated and undoes or interferes with the action of the PRC2 complex, to demethylate the Dmrt1 promoter at H3K27. Dmrt1, under the influence of its promoter, is then open for transcription and its expression frees the developmental trajectory toward a male fate, under the normal processes of reinforcement of male trajectory genes and antagonism of female trajectory genes.
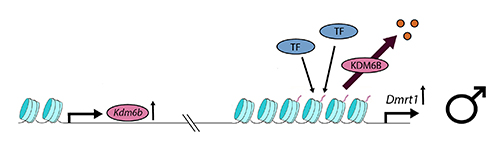
The story is not yet complete, because the process by which gene Kdm6b is thermosensitive is not yet clear. There is presumably some upstream thermosensitive element that governs expression of Kdm6b. So our understanding of the the process has been shifted one step back -- upstream regulators are yet to be discovered. The significance of the work by Ge et al. (2018) is that it has shifted dramatically the focus of inquiry from direct thermosensitivity of candidate sex determining genes to higher-order thermosensitive epigenetic processes that differentially release influential sex genes for expression. We are on the cusp of finally understanding the mechanisms by which temperature exerts its influence on sexual fate.
Remarkable stuff, but here matters get more complicated.
Work in our lab, reported by Deveson et al. (2017), uncovered temperature sensitive instances of intron retention during the transcription of Jumonji family genes, of which Kdm6b (=Jmjd3) is one. Kdm6b and Jarid2 each retain an intron, littered with stop codons, at low, male-producing temperatures in turtle. In a sense, this is an unwelcome complication, because possible loss of function of protein KDM6B at the very temperature at which it is upregulated by the processes outlined by Ge et al. could undermine its action in promoting male development. Kdm6b(IR) is massively upregulated at the low temperature of 26C, and so the component of normal KDM6B may be sufficient to deliver activity to Dmrt1. Alternatively, KDM6B(IR) may still retain relevant activity despite its truncation by the retention of intron 19. Future work will unravel this conundrum.
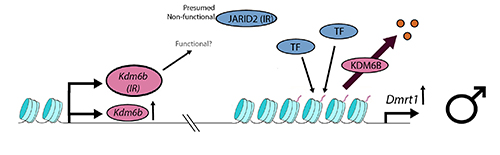
Deactivation of JARID2(IR) is more consistent with the interpretation reported by Ge et al.. Loss of function of JARID2(IR) means potentially that it is not facultatively available to form a complex with PRC2 to form PRC2.2 with ability to bind to the the promoter of Dmrt1. So at the lower male-producing temperatures, repressive trimethylation of the Dmrt1 promoter at H3K27 may be impeded, assisting the activation of Dmrt1.
These processes are occurring in other taxa, in the American Alligator (TSD) and the Central Bearded Dragon (temperature-induced sex reversal). The difference between them an the turtle is that the intron retention occurs at high female producing temperatures. What is the explanation for this?
A central role for these highly conserved and fundamental processes of chromatin modification leaves open the possibility that different sex genes can become enlisted to function as temperature-sensitive sex-determining genes, thus explaining the astonishing diversity of sex determination in reptiles. Perhaps in Alligator and Dragon, different sex-influential genes are the target of activation by demethylation which may explain the reverse pattern in comparison with turtles.
Clearly there is much to do and learn. The above scenarios derive in large part from work undertaken in mammals, and as such, should be considered as working hypotheses subject to confirmation or refutation by experiments using reptiles. Those experiments are underway, driven in large part by the impetus given by the outstanding work of Ge et al..
Further Reading
Aldiri, I., Vetter, M.L., 2012. PRC2 during vertebrate organogenesis: A complex in transition. Developmental Biology 367, 91–99.
Deveson, I.W., Holleley, C.E., Blackburn, J., Marshall Graves, J.A., Mattick, J.S., Waters, P.D., Georges, A., 2017. Differential intron retention in Jumonji chromatin modifier genes is implicated in reptile temperature-dependent sex determination. Science Advances 3, e1700731.
Ge, C., Ye, J., Weber, C., Sun, W., Zhang, H., Zhou, Y., Cai, C., Qian, G., Capel, B., 2018. The histone demethylase Kdm6b regulates temperature-dependent sex determination in a turtle species. Science.
Ge, C., Ye, J., Zhang, H., Zhang, Y., Sun, W., Sang, Y., Capel, B., Qian, G., 2017. Dmrt1 induces the male pathway in a turtle species with temperature-dependent sex determination. Development 144, 2222–2233.
Herpin, A., Schartl, M., 2011. Sex Determination: Switch and Suppress. Current Biology 21, R656–R659.
Marasca, F., Bodega, B., Orlando, V., 2018. How polycomb-mediated cell memory deals with a changing environment: Variations in PcG complexes and proteins assortment convey plasticity to epigenetic regulation as a response to environment. BioEssays 1700137.
Sawan, C., Herceg, Z., 2010. Histone Modifications and Cancer. Advances in Genetics 70, 57–85.