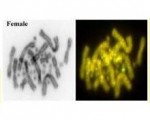
Genomics is currently undergoing a revolution brought about by the advent of high-throughput parallel sequencing and powerful cytogenetic techniques such as fluorescent in-situ hybridization (FISH) and comparative genomic hybridization (CGH). This has opened new opportunities to study non-model organisms, such as reptiles and amphibians, in ways that were previously restricted to human and mouse.
Of special interest to our lab is the comparative genomics of sex determination in reptiles. Sex determination has been a topic of speculation and rigorous inquiry since the time of Aristotle, and remains a hot topic today because of its intrinsic interest as a fundamental biological process, and because greater understanding brings benefit for human health. Our team at the University of Canberra is using frontier DNA technologies to probe the astonishingly diverse mechanisms of sex determination in reptiles.
As part of this research, we have collaborated with the Beijing Genome Institute (BGI, Shenzhen) to generate an annotated genome for the Australian central bearded dragon Pogona vitticeps. The annotated genome is complete, published in GigaScience, and a dense physical mapping of the genome is now also available through BMC Genomics. The resource includes a BAC library available commercially from Amplicon Express and a series of transcriptomes from a range of tissues.
The US National Center for Biotechnology Information (NCBI) has made the annotated genome sequence of Fabian, the ZZ dragon lizard, available to the international research community. The IAE made the genome available in 2013 through its local server, but we expect uptake to accellerate now that it is available through NCBI. Through the NCBI servier it is possible to BLAST sequence against the Pogona genome, pull down the sequences and annotation, and search for specific genes and regions.
To browse the genome, researchers will still have to access the GBrowser on the IAE server. This is soon to be upgraded to JBrowse with greater functionality and hosted on an Amazon WS. Watch this space.
Further information on how to access the genome,associated transcriptomes and publications click here.